Pandas in Python: Use with Hydrological Time Series
I recently had some time series analysis to do, and I decided to do this work in Pandas. Particualarly, I had to deal with many timeseries, stretching from different startpoints in time to different endpoints in time. Each timeseries was available as an ASCII file. The data were daily data. The time information was given in three columns, each representing year, month, and day, respectively.

Image: http://flic.kr/p/9gK3ZH
I found these two posts by Randal S. Olson very useful resources:
- Using pandas DataFrames to process data from multiple replicate runs in Python (link)
- Statistical analysis made easy in Python with SciPy and pandas DataFrames (link)
Here is a cookbook style layout of what I did:
The following steps show how easy it was to deal with the data 1. Read the input data
cur_sim_ts = pa.io.parsers.read_table(os.path.join(data_path, 'test', result)
, header=None
, sep='\s*'
, parse_dates=[[0,1, 2070]]
, names=['year','month', 'day', result[:-4]]
, index_col=[0]
)
where `’\s*’ means any whitespace. The dates were given in three colums, one for year, month, and day. Can it get simpler and nicer than that?
It is possible to repeat 1 multiple time, each time extending the pandas data_frame. Unfortunately, this looks a little ugly still, but works
if counter_cur_file > 0: final_df = final_df.combine_first(cur_sim_ts) else: final_df = cur_sim_ts.copy() / 10.0In the else part, the Pandas data_frame is initialised. It so happens that this and only this series of the loop has to be divided by ten. In all other cases the time_series that was read in step 1 is “appended” (or rather combined with) the the previously initialized data_frame. The wicked thing is that each time_series is put at the propper “place” in time within the data_frame. Dates are real dates. This is beautiful, but I had to be a little carfule with the data I had at hand, in which every month has 30 days.
As soon as this data_frame is constructed, things are easy, for example
plotting, particularly plotting only a certain time-interval of the data.
final_df['2070-01-01':'2100-12-30'].plot(ylim=[-10,45])saving the data_frame
final_df.save(os.path.join(out_path, pickle_filename))
For me it was of particular interest to find out, how many consecutive dry and wet days there are in each time series. I introduced a threshold of precipitation. If the daily amount of precipitation is above that threshold, this day is considered to be “wet”, else it’s considered to be “dry”. I wanted to count the number of consecutive dry and wet days, and remember them for a time series. This is the purpose of the function below. It is coded a little bute force. Still I was surprised, that it performed reasonably well. If anybody has a better idea, please let me know. Maybe it can be of use for other Pandas users. Note: a time_series in the Pandas world is obtained by looping over a data_frame
def dry_wet_spells(ts, threshold): """ returns the duration of spells below and above threshold input ----- ts a pandas timeseries threshold threshold below and above which dates are counted output ------ ntot_ts total number of measurements in ts n_lt_threshold number of measurements below threshold storage_n_cons_days array that stores the lengths of sequences storage_n_cons_days[0] for dry days storage_n_cons_days[1] for wet days """ # total number in ts ntot_ts = ts[~ ts.isnull()].count() # number lt threshold n_lt_threshold = ts[ts <= threshold].count() # type_day = 0 # dry # type_day = 1 # wet # initialisierung: was ist der erste Tag type_prev_day = 0 storage_n_cons_days = [[],[]] n_cons_days = 0 for cur_day in ts[~ ts.isnull()]: # current day is dry if cur_day <= threshold: type_cur_day = 0 if type_cur_day == type_prev_day: n_cons_days += 1 else: storage_n_cons_days[1].append(n_cons_days) n_cons_days = 1 type_prev_day = type_cur_day else: type_cur_day = 1 if type_cur_day == type_prev_day: n_cons_days += 1 else: storage_n_cons_days[0].append(n_cons_days) n_cons_days = 1 type_prev_day = type_cur_day return ntot_ts, n_lt_threshold, storage_n_cons_days- With all of this, I can produce histograms like this:
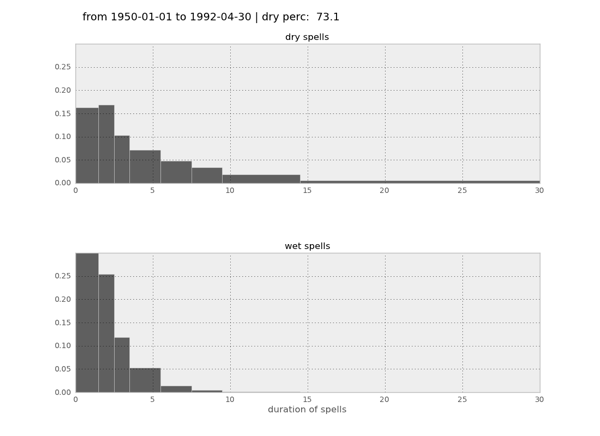
Hello, I stumbled on your page when I was googling how to count consecutive dry days. I am brand new to Python, and I am currently practicing with daily precipitation data (60 or so years worth of data). So far, I have managed to separate year, month, and day into three separate columns. I need to count how many episodes there is with consecutive dry days (0 mm of precipitation). How would I go about it? I did try your codes, but I think I am doing something wrong as it doesn’t work with my data. Thank you for your time!
Jocelyn
1 Mar 18 at 10:56 pm
Dear Jocelyn, the code above takes daily input data. The second argument,
thresholdacts as an indicator, everything below counts as dry, everything above as wet. You could setthresholdto zero. However, I would encourage you to use some value slightly larger than zero, because a precipitation of 0.1mm does not contribute to any flow.How does your data look like – could you be more specific?
Claus
2 Mar 18 at 10:41 am
Hello! I have a data frame with ‘Year’, ‘Month’, ‘Day’, ‘Precipitation’, ‘MinimumTemp’, and ‘MaximumTemperature columns. I have daily precipitation data ranging from 1950 to 2012. For now, I am just going to practice with one year of data (1950). I want Python to count the number of episodes with consecutive dry days within that year. Hopefully that was specific. Thank you!!
Jocelyn
3 Mar 18 at 1:34 am
If I understand you correctly, you need to provide the column with
precipitationto your script and it will return to you the histogram of the duration of dry spells – isn’t that what you want?Claus
5 Mar 18 at 9:38 am
Hello so I have a time series of precipitation. I want to find the number of dry months below threshold of -1 (data is SPI). if I have more two months below this threshold then I count. it consecutive dry and wet days. this code doesnt help me in this regards,
thanks
Staph
28 Apr 19 at 1:24 pm
Hi Claus,
I have used your code to try and count the number of days per year that a fire weather index is above or below a set threshold. For the most part, the code worked well, but I have noticed that there are some errors in the total counts. For example, (total days meeting or exceeding threshold) + (total days below threshold) does not equal the total number of days in a year. And similarly, when looking at the graphed data, I count x number of distinct periods (above or below threshold), but the results of the code indicate there are (x-1) distinct periods.
I have essentially used your exact code, so I’m wondering if you may have any insight into this problem or have encountered similar problems in your own program.
Thanks!
Ben
29 Jun 19 at 12:16 am
Dear Ben, nice to hear that it works generally well. Related to your first question: I’m afraid I have never tried this for periods longer than a year, as there is nothing implemented in relation to number of days in months/years. Related to your second question: I am uncertain what exactly you are plotting, but there is always the danger of an index/counting error.
Claus
29 Jun 19 at 4:00 pm